Published: Modeling the evolutionary dynamics of clonal hematopoiesis

EDIT lab has recently published a Perspective paper in Nature Genetics, entitled “Modeling the evolutionary dynamics of clonal hematopoiesis.”
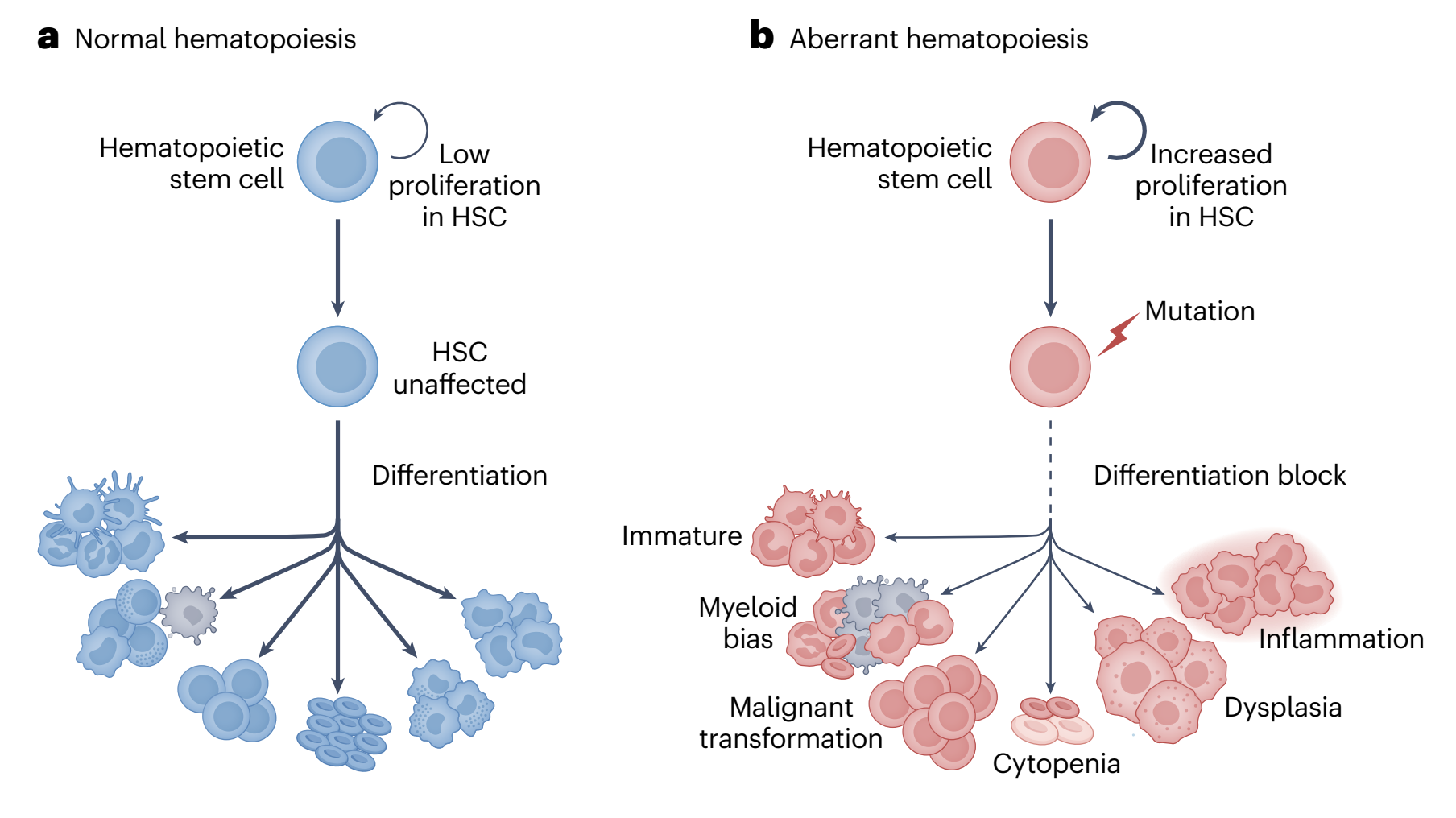
In this paper, we integrate and formalize mathematical and computational modeling frameworks to quantify the evolutionary behavior of hematopoietic stem cell (HSC) clones over the human lifespan. Clonal hematopoiesis (CH) is not merely a descriptive genomic phenomenon; it is a dynamical system governed by mutation, selection, drift, stem cell pool constraints, inflammation, aging, and therapy-induced perturbations. We compare deterministic and stochastic modeling paradigms, including exponential and logistic models, Moran and extended Moran processes, branching processes, coalescent theory, and site-frequency spectrum (SFS) inference, and evaluate the biological validity of their underlying assumptions in homeostatic versus treatment-perturbed hematopoiesis. Importantly, we emphasize that fitness is a context-dependent and inherently mathematical quantity that cannot be inferred qualitatively from variant allele frequency (VAF) trajectories alone. This work advances a constrained, evolution-informed framework that shifts CH from descriptive genomics to predictive, patient-specific modeling, linking molecular-scale mutations to clinical risk stratification.




Comments